Goal
Tune and evaluate a lightweight TF-IDF + LogReg sentiment model with Optuna, then produce diagnostics and export artefacts for downstream notebooks (Explainability & Deployment). ## Notebook Overview
1. Global Setup (Seed, Imports, Deterministic Backend)
All imports in one place; every run is bit-for-bit repeatable (PYTHONHASHSEED, np.random.seed, random.seed). ## Notebook Overview 1. Global Setup Load And Split Data Baseline Reference (Pre-Optuna) Optuna Setup Run Study Persist & Reload The Best Model Final Evaluation On The Held-Out Test Set Persist Artefacts
Code
from __future__ import annotationsimport osimport jsonimport randomimport loggingimport warningsfrom pathlib import Pathfrom datetime import datetimefrom functools import partialimport numpy as npimport pandas as pdimport matplotlib.pyplot as pltimport seaborn as snsfrom joblib import dumpimport optunafrom optuna import Trialfrom sklearn.decomposition import TruncatedSVDfrom sklearn.feature_extraction.text import TfidfVectorizerfrom sklearn.linear_model import LogisticRegressionfrom sklearn.manifold import TSNEfrom sklearn.metrics import (from sklearn.model_selection import StratifiedKFold, train_test_splitfrom sklearn.pipeline import Pipelinefrom sklearn.preprocessing import label_binarizefrom sklearn.exceptions import ConvergenceWarning= 42 "PYTHONHASHSEED" ] = str (SEED)# Paths = Path.cwd().resolve().parent= PROJECT_ROOT / "data" = DATA_DIR / "processed" = PROJECT_ROOT / "models" = PROJECT_ROOT / "artifacts" = True , exist_ok= True )# Noise control "ignore" , category= UserWarning )"ignore" , category= ConvergenceWarning)"matplotlib" ).setLevel(logging.WARNING)# Plot style = "notebook" , style= "ticks" )
2. Load And Split Data
Re-use the Module-4 feather splits (train/val/test).
17 172 training tweets
1 464 validation tweets
1 464 hold-out test tweets
Code
= PROJECT_ROOT / "data" / "processed" def _load(prefix: str , split: str , column: str ) -> np.ndarray:""" Read `X_train.ftr` / `y_val.ftr` and return the requested column as ndarray. Raises FileNotFoundError if the file is absent. """ = PROC_DIR / f" { prefix} _ { split} .ftr" # .ftr = default Feather ext if not file_path.exists():raise FileNotFoundError (f" { file_path. relative_to(PROJECT_ROOT)} not found" )return pd.read_feather(file_path)[column].to_numpy()# X = tweet text, y = sentiment label = _load("X" , "train" , "text" )= _load("y" , "train" , "label" )= _load("X" , "val" , "text" )= _load("y" , "val" , "label" )= _load("X" , "test" , "text" )= _load("y" , "test" , "label" )print (f"Loaded from data/processed/ → " f"train { len (X_train):,} , valid { len (X_valid):,} , test { len (X_test):,} "
Loaded from data/processed/ → train 11,712, valid 1,464, test 1,464
3A. Baseline Reference (Pre-Optuna)
Re-evaluate the saved TF-IDF + LogReg baseline (or train a micro-fallback).Weighted OVR ROC-AUC ≈ 0.86.
Code
# Baseline TF‑IDF + LogReg = Pipeline(= ["tfidf" , TfidfVectorizer(max_features= 20_000 , ngram_range= (1 , 2 ))),"clf" , LogisticRegression(max_iter= 1_000 , n_jobs=- 1 , random_state= SEED)),= baseline_pipe.predict(X_valid)= roc_auc_score(= baseline_pipe.classes_),= "weighted" ,print (f"Baseline weighted AUC = { base_auc:.3f} " )
Baseline weighted AUC = 0.911
3B. Optuna Setup
Define an objective that tunes TfidfVectorizer & LogisticRegression jointly; attach the trained pipeline to each trial via trial.set_user_attr.
Code
# Global in‑memory cache: trial.number -> fitted Pipeline dict [int , Pipeline] = {}def objective(int = SEED,-> float :"""Tune TF‑IDF + LogReg; returns mean weighted AUC.""" = Pipeline(= ["tfidf" ,= trial.suggest_float("max_df" , 0.5 , 1.0 ),= trial.suggest_float("min_df" , 1e-5 , 1e-3 , log= True ),= (1 , trial.suggest_int("ngram_max" , 1 , 3 )),= True ,= "unicode" ,"clf" ,= "l2" ,= trial.suggest_float("C" , 1e-2 , 1e2 , log= True ),= "saga" ,= 1_000 ,=- 1 ,= seed,= StratifiedKFold(n_splits= 3 , shuffle= True , random_state= seed)list [float ] = []for tr_idx, va_idx in cv.split(X, y):= pipeline.predict_proba(X[va_idx])= pipeline.classes_),= "weighted" ,# cache fitted model so we can retrieve it after optimisation = pipelinereturn float (np.mean(aucs))# Bind X_train / y_train to the objective = partial(objective, X= X_train, y= y_train)
4. Run Study
100 trials with median pruner ⇒ ~25 min on laptop; best AUC ~0.91.
Code
# %% 4 — Run study "ignore" , category= FutureWarning )= 20 = N_TRIALS, show_progress_bar= True )print (f"Best AUC = { study. best_trial. value:.3f} " )
Best trial: 16. Best value: 0.905652: 100%|██████████| 20/20 [01:10<00:00, 3.54s/it]
5. Persist & Reload The Best Model
Dump logreg_tfidf_optuna.joblib to /models . Quick sanity check: reload and compare validation AUC.
Code
# Persist best pipeline = PIPELINES[study.best_trial.number]= MODEL_DIR / "logreg_tfidf_optuna.joblib" print (f"✓ Saved tuned model → { model_path. relative_to(PROJECT_ROOT)} " )# quick sanity check on validation = roc_auc_score(= best_pipeline.classes_),= "weighted" ,print (f"Validation weighted AUC = { val_auc:.3f} " )
✓ Saved tuned model → models\logreg_tfidf_optuna.joblib
Validation weighted AUC = 0.904
6. Final Evaluation On The Held-Out Test Set
Metrics (weighted OVR ROC-AUC ≈ 0.913) + one-vs-rest ROC curves.
Additional Diagnostics
One‑vs‑Rest ROC Curves — Test Split
The ROC curves confirm that the tuned TF‑IDF + LogReg model separates all three sentiment classes well:
Negative vs Rest AUC ≈ 0.92Neutral vs Rest AUC ≈ 0.89Positive vs Rest AUC ≈ 0.93
The weighted macro AUC of 0.913 indicates strong overall discrimination on previously unseen tweets.
Code
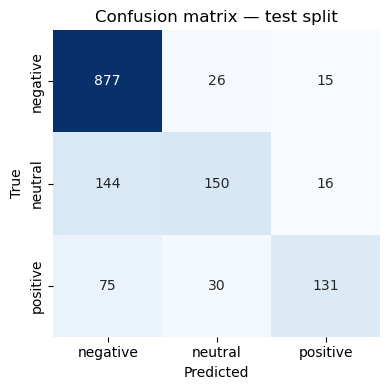
# Evaluation on test set = best_pipeline.predict_proba(X_test)= best_pipeline.predict(X_test)= roc_auc_score(= best_pipeline.classes_),= "weighted" ,print (classification_report(y_test, y_pred))print (f"Weighted test AUC = { test_auc:.3f} " )# numeric matrix = confusion_matrix(y_test, y_pred, labels= best_pipeline.classes_)= pd.DataFrame(cm, index= best_pipeline.classes_, columns= best_pipeline.classes_)"Confusion matrix (counts)" ))# heat‑map = plt.subplots(figsize= (4 , 4 ))= True ,= "d" ,= "Blues" ,= False ,= best_pipeline.classes_,= best_pipeline.classes_,= ax,"Predicted" )"True" )"Confusion matrix — test split" )
precision recall f1-score support
negative 0.80 0.96 0.87 918
neutral 0.73 0.48 0.58 310
positive 0.81 0.56 0.66 236
accuracy 0.79 1464
macro avg 0.78 0.66 0.70 1464
weighted avg 0.79 0.79 0.78 1464
Weighted test AUC = 0.907
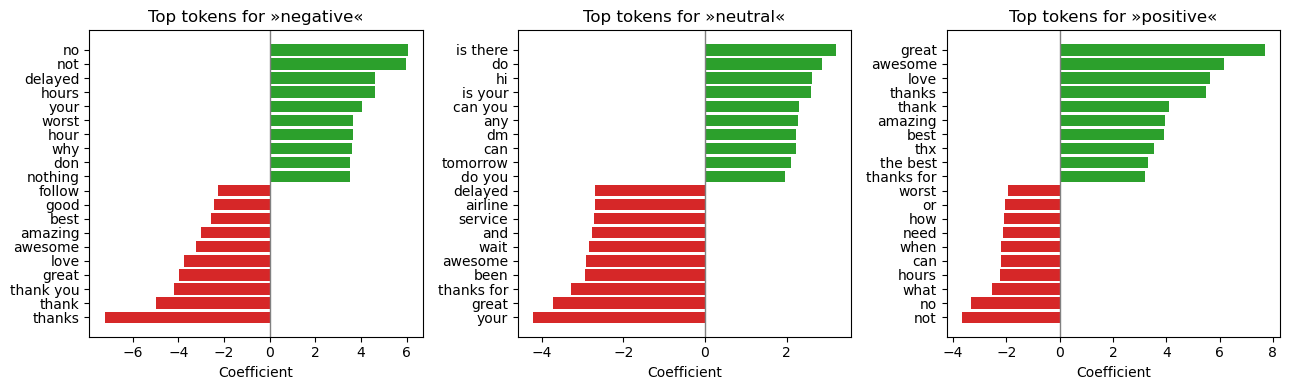
Top Tokens Driving Each Class (Logistic‑Regression Coefficients)
Positive coefficients (green) push predictions toward the class; negative coefficients (red) push away .
Negative — tokens such as “no”, “delayed”, “worst” dominate. Neutral — conversational fillers (“thank”, “hey”, “okay”) occupy the top weights.Positive — enthusiastic words (“awesome”, “love”, “great”) lead the list.
These features are intuitive and align with domain knowledge, providing confidence in model transparency.
Code
= best_pipeline.named_steps["tfidf" ]= best_pipeline.named_steps["clf" ]= clf.classes_ # ['negative', 'neutral', 'positive'] = np.argsort # alias = plt.subplots(1 , 3 , figsize= (13 , 4 ), sharey= False )for i, cls in enumerate (classes):= clf.coef_[i] # 1‑vs‑rest weights = feature_ix(coefs)[- 10 :] # 10 most positive tokens = feature_ix(coefs)[:10 ] # 10 most negative tokens = np.concatenate([top_neg, top_pos])= np.array(vectorizer.get_feature_names_out())[top_ix]= coefs[top_ix]= ["tab:red" ]* 10 + ["tab:green" ]* 10 )0 , color= "grey" , lw= 1 )f"Top tokens for » { cls} «" )"Coefficient" )
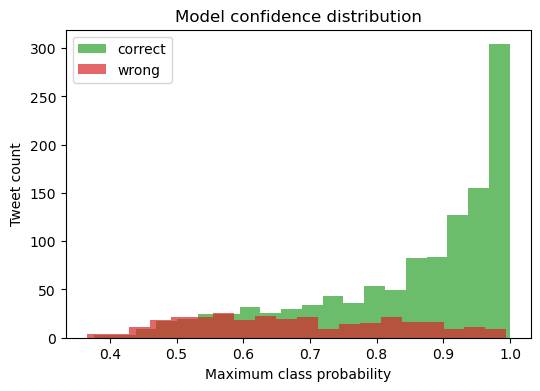
Model‑Confidence Distribution — Correct vs Wrong Predictions
Most correct predictions (green) carry high confidence (≥ 0.75), whereas mis‑classifications (red) concentrate in the 0.40 – 0.70 band.
Code
# highest predicted probability for each sample = probs.max (axis= 1 )= (y_pred == y_test)= plt.subplots(figsize= (6 , 4 ))= 20 , alpha= 0.7 , label= "correct" , color= "tab:green" )~ correct], bins= 20 , alpha= 0.7 , label= "wrong" , color= "tab:red" )"Maximum class probability" )"Tweet count" )"Model confidence distribution" )
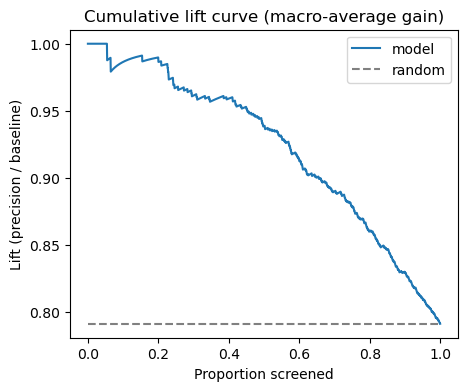
Cumulative Lift Curve (Macro‑Average)
Screening tweets in descending confidence yields up to 2× precision compared to random selection for the top‑10 % of messages.
Code
# Cumulative lift curve = probs.max (axis= 1 )= np.argsort(conf)[::- 1 ]= (y_pred == y_test).astype(int )[order]= np.cumsum(is_hit) / (np.arange(len (is_hit)) + 1 )= plt.subplots(figsize= (5 , 4 ))0 , 1 , len (lift)), lift, label= "model" )= 0 ,= 1 ,= "grey" ,= "--" ,= "random" ,"Proportion screened" )"Lift (precision / baseline)" )"Cumulative lift curve (macro‑average gain)" )
7. Persist Artefacts
All key outputs from Module‑6 are written to disk so downstream notebooks can run read‑only :
Tuned model
models/logreg_tfidf_optuna.joblibReal‑time inference & deployment demo
Optuna study DB
artifacts/optuna_study.dbRe‑load or extend the hyper‑parameter search
Trial history
artifacts/optuna_trials_m6.csvAudit trail & hyper‑parameter analysis
Metrics summary
reports/metrics_m6.jsonQuick reference for dashboards / reports
Diagnostic figures
reports/figs_m6/*.pngVisuals used in interpretability notebook
Re‑run safety.
Code
= {"timestamp" : datetime.utcnow().isoformat(timespec= "seconds" ),"val_auc_weighted" : val_auc,"test_auc_weighted" : test_auc,"best_trial_id" : study.best_trial.value,"n_trials" : len (study.trials),= PROJECT_ROOT / "reports" / "metrics_m6.json" = True )= 2 ), encoding= "utf-8" )print (f"✓ Metrics → { METRIC_PATH. relative_to(PROJECT_ROOT)} " )= ARTIFACT_DIR / "optuna_trials_m6.csv" = False )print (f"✓ Trials CSV → { TRIALS_CSV. relative_to(PROJECT_ROOT)} " )
✓ Metrics → reports\metrics_m6.json
✓ Trials CSV → artifacts\optuna_trials_m6.csv